Msg1 = "Adjust brightness and contrast and press \"OK\"." Manually adjust B+C if needed, if not then just press OKĭialog.addChoice("Type:", newArray("YEP", "NOPE")) REMEMBER to change the file names between samples 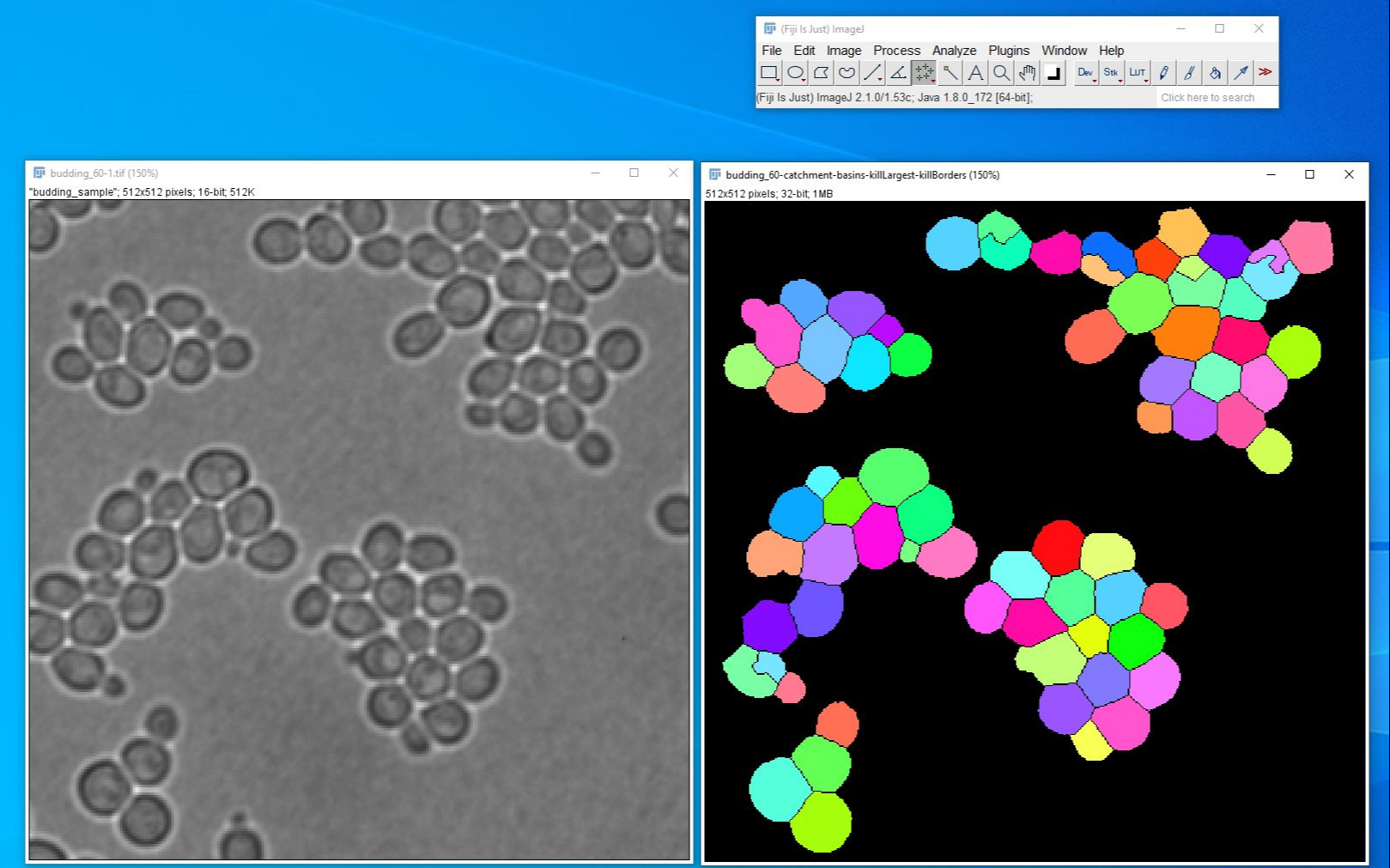
Running an automated save, the original stack (unadjusted B+C) will be saved as a TIFF in a directory of your choosing. Run("Make Composite", "display=Composite") Before these steps I select my cell and rotate the stack etc. That way I can make sure that the DIC is completely in focus and at a good contrast but I can still see the complexity of the max projection in the colour channels.Ĭan anyone help me with this? I’ve pasted the middle to end of my script below. Then at the end of the panel I want an overlay with the max projected colour channels on top of a DIC image from a selected z-slice. What I want is my colour channels (magenta, green, DAPI) each max projected and displayed as single images in a panel so that we can see each channel clearly. The DIC images in the max projections look awful so I need to adapt my script slightly. I’ve developed a basic script which allows me to do this with single slices or by max projecting the whole stack. I have 3 (sometimes 4) distinct channels, including DAPI and DIC and want to make a montage to display my data as a panel of single channels followed by an overlay with the colours+DIC. I use z-stacks (60 individual slices) to analyse my cell biology data. 
I’m hoping that someone can help me with adjusting my FIJI script to combining both max projected z-stacks and single z-slices into a single montage. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
